博文
hLife Collection | Viruses (Part Ⅱ)
|
1. ACE2-using coronaviruses: A global concern
通信作者:刘科芳、高福
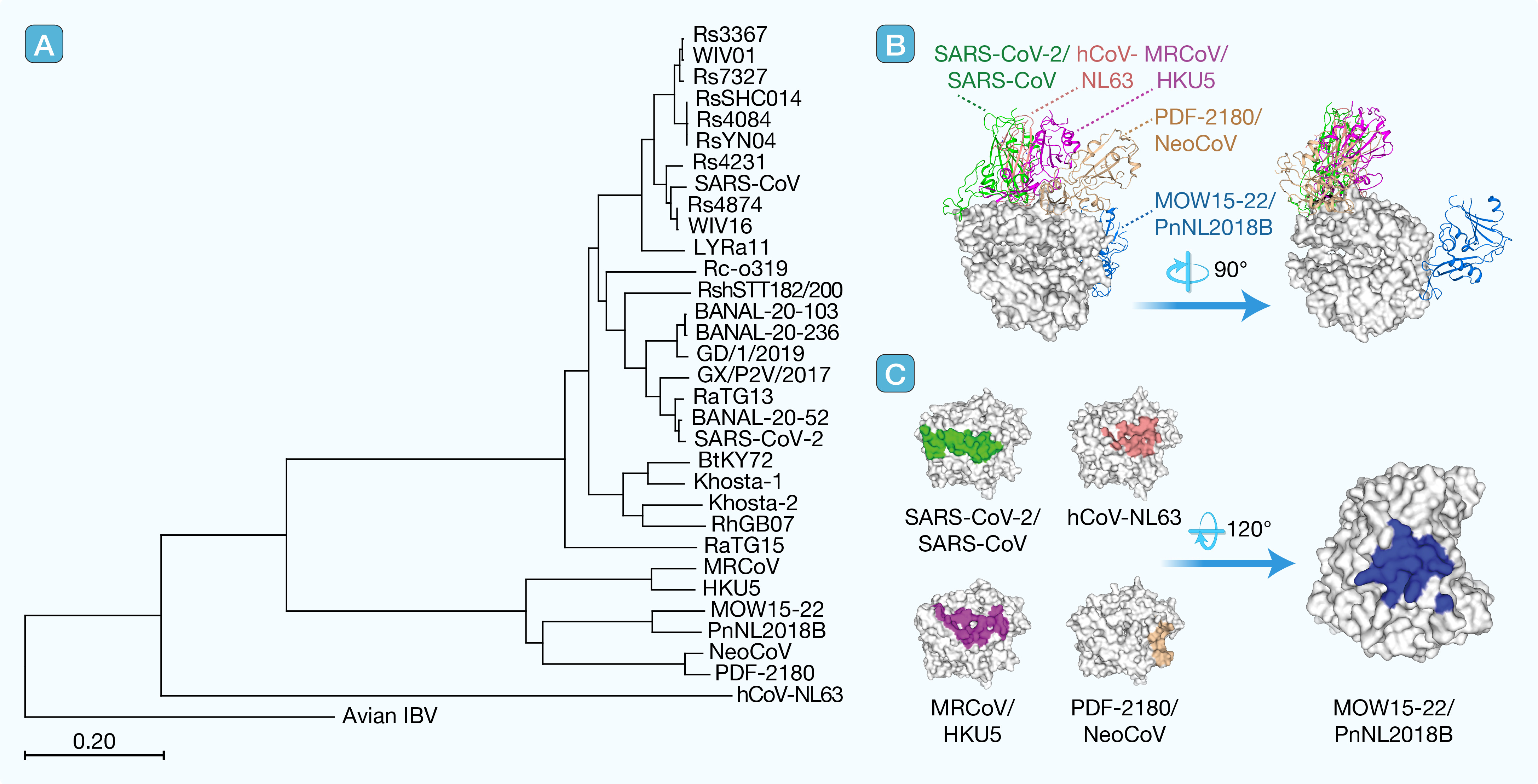
引用:
Xu Z, Lobato AC, Liu K, et al. ACE2-using coronaviruses: A global concern. hLife 2025; 3: 615–617.
hLife | 中国科学院高福院士团队开发BAADesign:赋能埃特司韦单抗破解新冠Omicron变异株挑战
通信作者:杨梦苏、高福
 引用:
引用:Su C, He J, Xie Y, et al. Enabling the immune escaped etesevimab fully-armed against SARS-CoV-2 Omicron subvariants including KP.2. hLife 2025; 3: 132–145.
通信作者:张洪春、毕玉海
 引用:
引用:Li J, Huan Y, Xia Q, et al. H3N2 influenzavirus characteristics in China (2019–2022): Genetic, antigenic, and infection dynamics during the COVID-19 pandemic. hLife 2025; 3: 146–158.
4.Modeling viral evolution: A novel SIRSVIDE framework with application to SARS-CoV-2 dynamics
hLife | 陆剑研究团队研发SIRSVIDE模型解析病毒进化动态
通信作者:陆剑
本研究研发了SIRSVIDE( Susceptible-Infected-Recovered-Susceptible-Variation-Immune Decay-Immune Escape )的新型计算模型,用于模拟病毒种群的传播和进化动态。
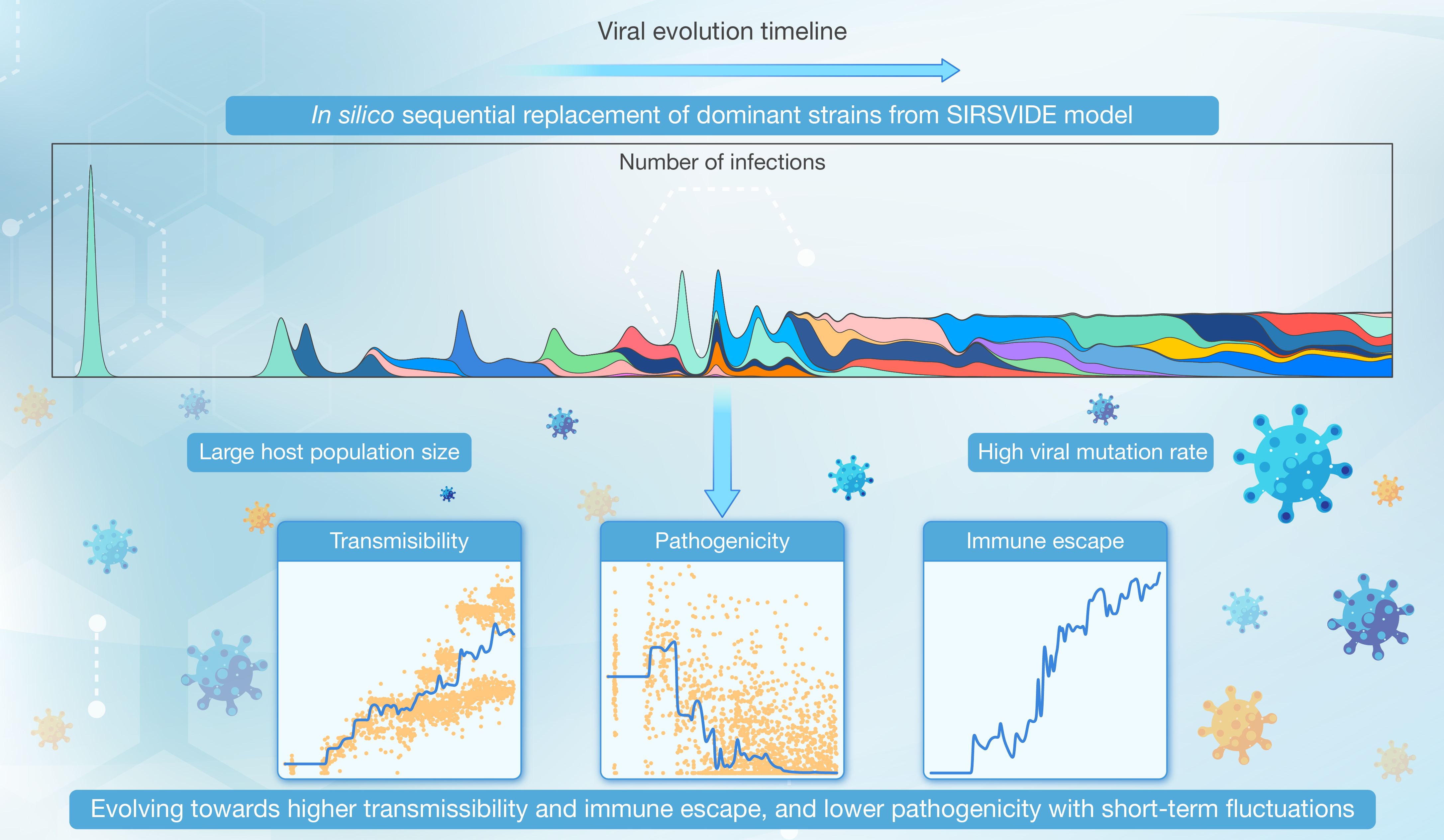 引用:
引用:Jin K, Tang X, Qian Z, et al. Modeling viral evolution: A novel SIRSVIDE framework with application to SARS-CoV-2 dynamics. hLife 2024; 2: 227–245.
通信作者:Jana Shen、Rolf Hilgenfeld
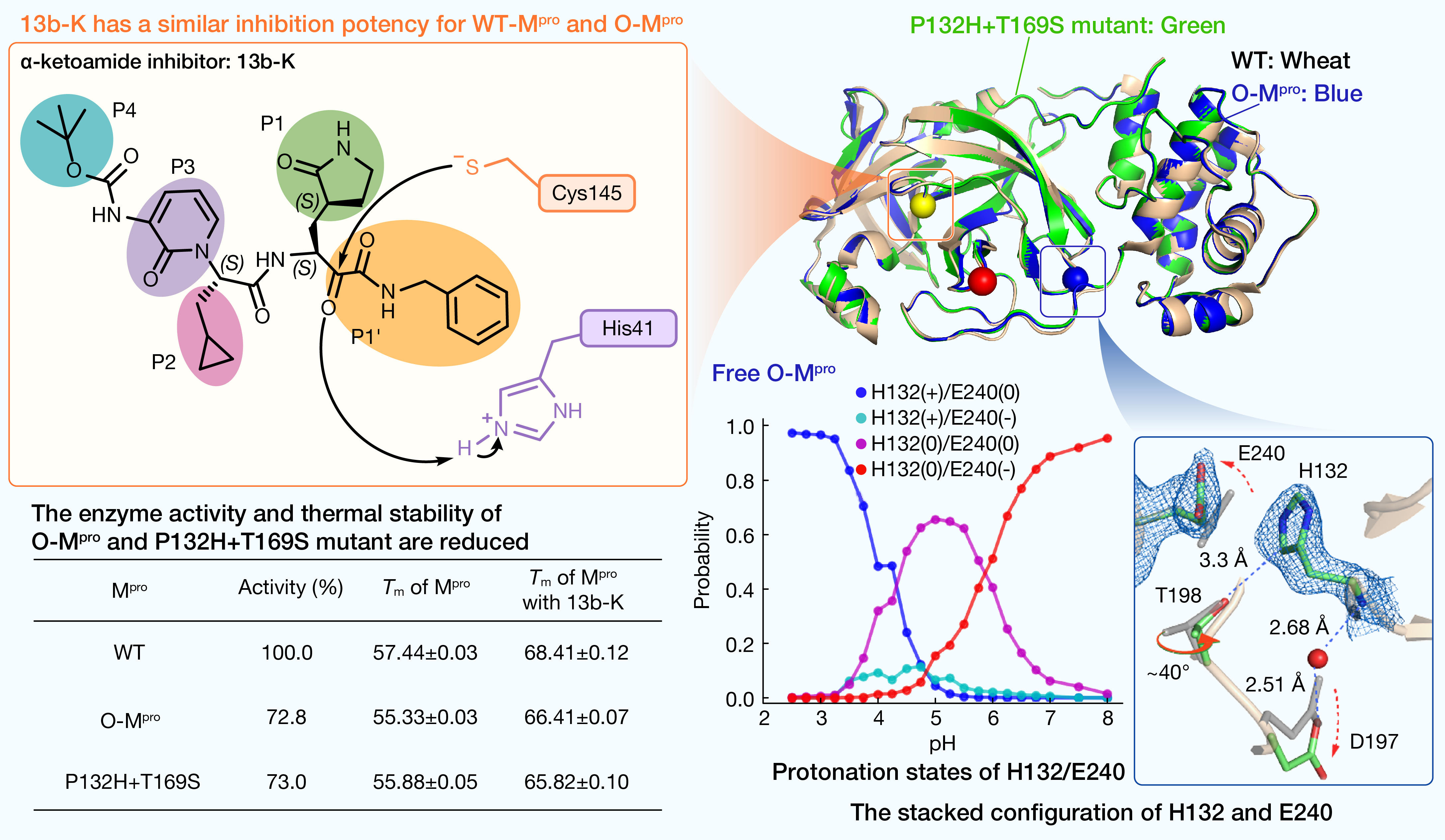 引用:
引用:Ibrahim M, Sun X, Martins de Oliveira V, et al. Why is the Omicron main protease of SARS-CoV-2 less stable than its wild-type counterpart? A crystallographic, biophysical, and theoretical study. hLife 2024; 2: 419–433.
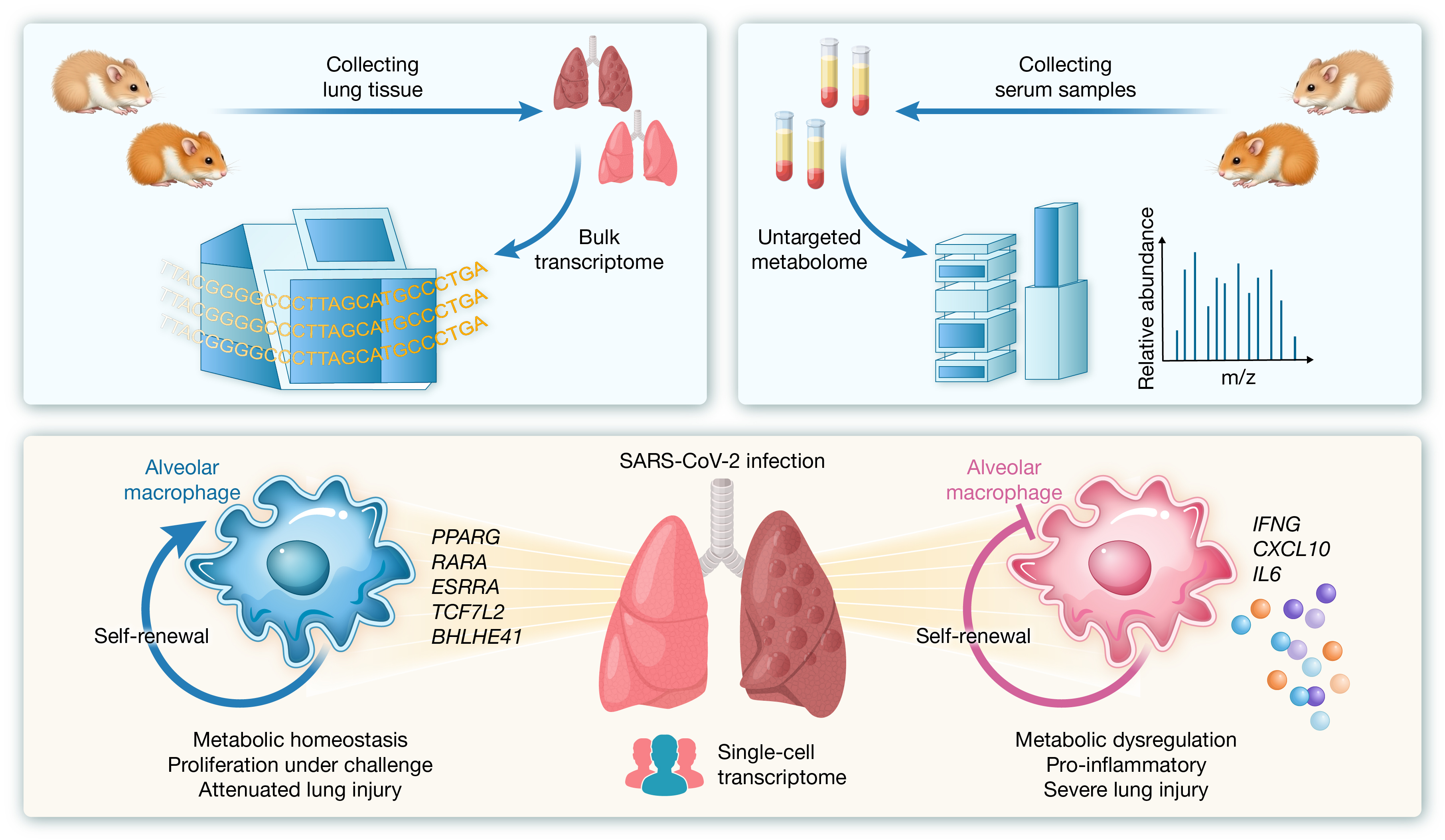
6. Prophylactic and therapeutic itaconate treatment alleviates COVID-19-associated lung injury
通信作者:林树海、张天英、夏宁邵
 引用:
引用:Jiang Y, Yang P, Yao B, et al. Prophylactic and therapeutic itaconate treatment alleviates COVID-19-associated lung injury. hLife 2025; 3: 551–564.
7. Selection and engineering of broad-spectrum antiviral affibody peptides against SARS-CoV-2 variants
通信作者:Christopher John Hipolito、齐建勋、施一
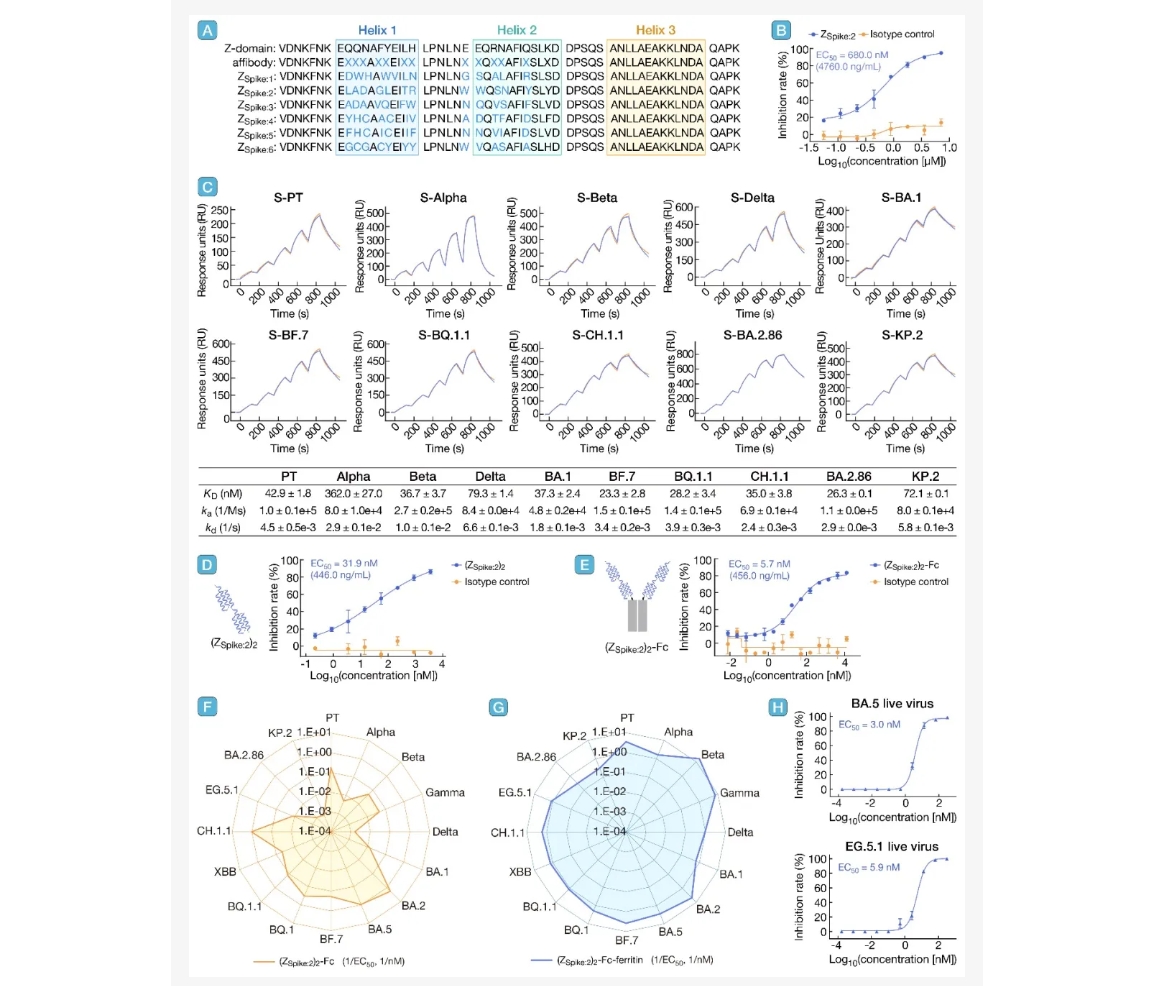 引用:
引用:Yang J, Wang M, Chen Z, et al. Selection and engineering of broad-spectrum antiviral affibody peptides against SARS-CoV-2 variants. hLife 2025; 3: 448–451.
通信作者:夏朋延
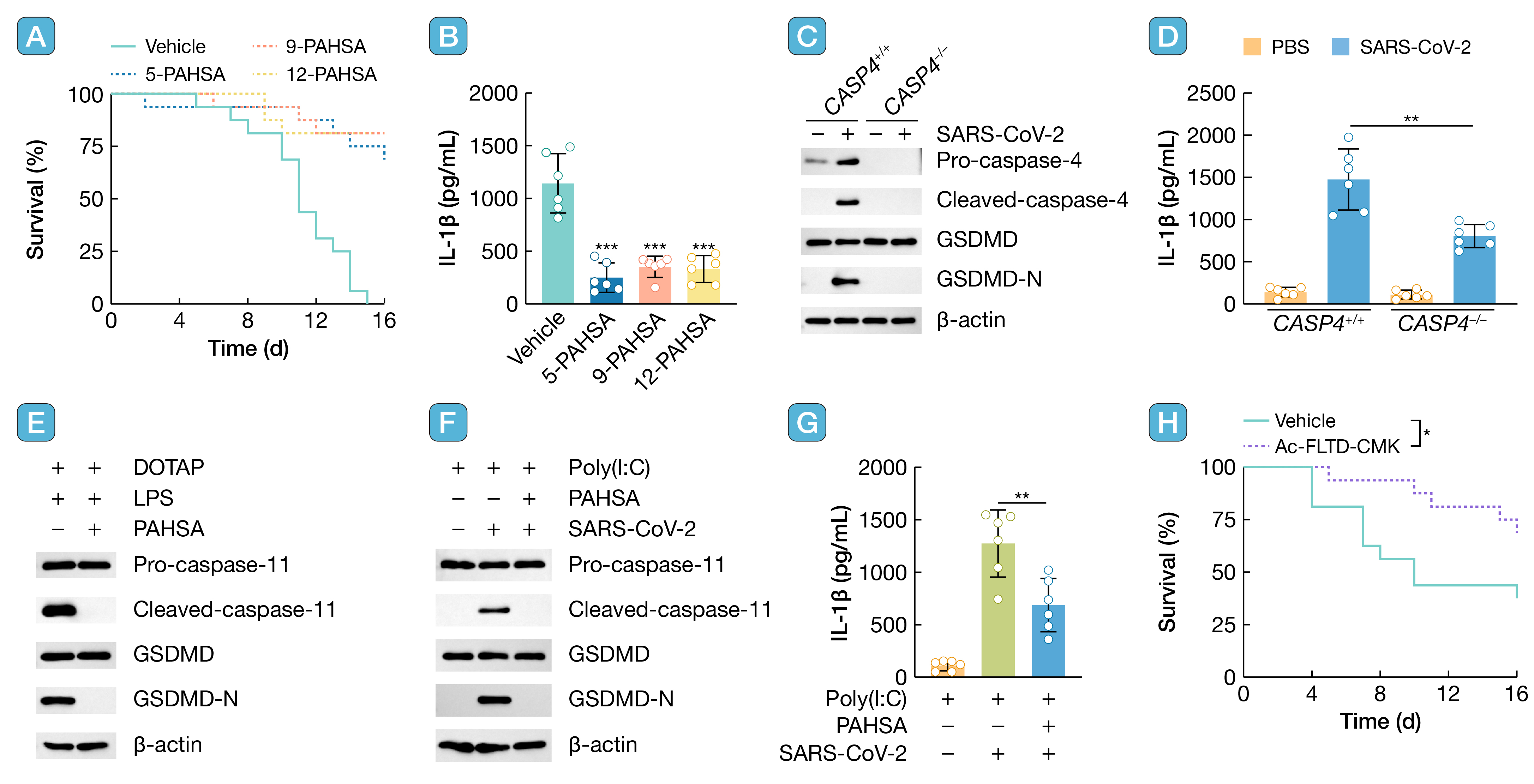 引用:
引用:Kong C, Li Y, Qian Y, et al. Palmitic acid esters of hydroxy stearic acids suppress SARS-CoV-2 infection through inhibiting the non-canonical inflammasome. hLife 2025; 3: 452–454.
通信作者:白崇智、苏超、韩鹏程
A novel pangolin-origin Middle East respiratory syndrome coronavirus-like (MERS-like coronavirus), Manis javanica HKU4-related coronavirus 1 (MjHKU4r-CoV-1), has recently been identified. This virus utilizes dipeptidyl peptidase 4 (DPP4 or CD26) as its entry receptor and exhibits high infectivity in human cells and organs. However, no therapeutic agents have been developed to inhibit a potential outbreak of MjHKU4r-CoV-1. In this study, we determined the Cryo-electron microscopy (Cryo-EM) structure of the MjHKU4r-CoV-1 receptor-binding domain (RBD) in complex with human CD26 (hCD26), revealing the detailed interactions at the virus-receptor interface. Based on these structural insights, we designed an hCD26 mutant (hCD26mu) with enhanced binding affinity for the MjHKU4r-CoV-1 RBD and demonstrated its potent neutralizing activity against MjHKU4r-CoV-1 pseudovirus. When fused to ferritin as a nanoparticle, the hCD26mu exhibited increased neutralization potency against both MjHKU4r-CoV-1 and MERS-CoV. These findings highlight the potential of the optimized hCD26mu as a therapeutic candidate for preventing MjHKU4r-CoV-1 infection.
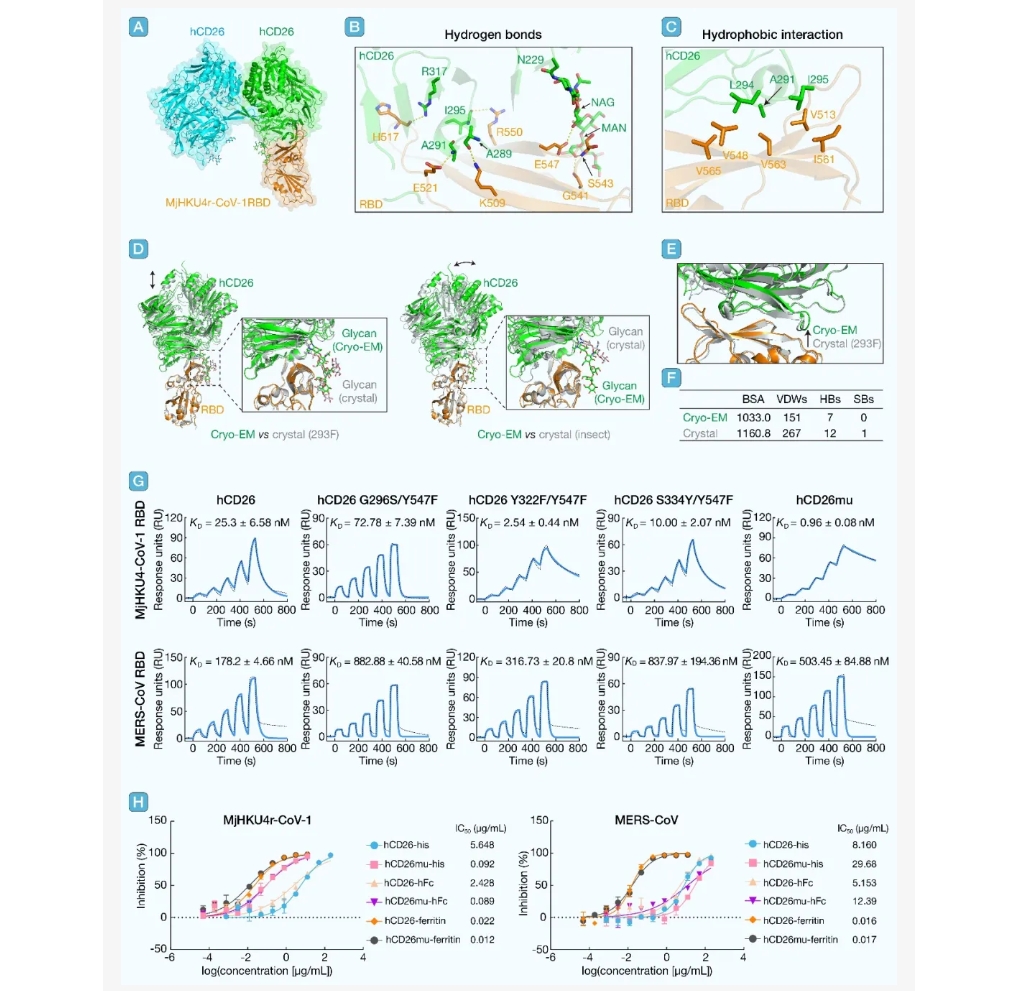 引用:
引用:Zhang Y, Tian M, Han Y, et al. Rational design of human CD26 receptor for a strong neutralizing ability against MjHKU4r-CoV-1 and MERS-CoV. hLife 2025; 3: 565–568.
通信作者:陈蓉,徐颖,张水军
This study presents the high-resolution structure of HCoV-HKU1A RBD bound to human TMPRSS2 receptor, which reveals the structural basis for the specificity of receptor binding by HCoV-HKU1. The interface between HCoV-HKU1 RBD and receptor also provides information for the development of vaccines and antivirals against HCoV-HKU1.
 引用:
引用:Wang W, Guan J, Ren M, et al. High-resolution crystal structure of human coronavirus HKU1 receptor binding domain bound to TMPRSS2 receptor. hLife 2024; 2: 653–657.
https://blog.sciencenet.cn/blog-3552961-1526293.html
上一篇:hLife Collection | AI for health science
下一篇:hLife Collection | Viruses (Part Ⅰ)