博文
Nature:2011年背靠背发表的植物感知氧气浓度分子机制
|||
Oxygen sensing in plants is mediated by an N-end rule pathway for protein destabilization
First author: Francesco Licausi; Affiliations: Max Planck Institute of Molecular Plant Physiology (马克斯·普朗克分子植物生理研究所): Potsdam-Golm, Germany
Corresponding author: Joost T. van Dongen
The majority of eukaryotic organisms rely on molecular oxygen for respiratory energy production. When the supply of oxygen is compromised, a variety of acclimation responses are activated to reduce the detrimental effects of energy depletion. Various oxygen-sensing mechanisms have been described that are thought to trigger these responses, but they each seem to be kingdom specific and no sensing mechanism has been identified in plants until now. Here we show that one branch of the ubiquitin-dependent N-end rule pathway for protein degradation, which is active in both mammals and plants, functions as an oxygen-sensing mechanism in Arabidopsis thaliana. We identified a conserved amino-terminal amino acid sequence of the ethylene response factor (ERF)-transcription factor RAP2.12 to be dedicated to an oxygen-dependent sequence of post-translational modifications, which ultimately lead to degradation of RAP2.12 under aerobic conditions. When the oxygen concentration is low—as during flooding—RAP2.12 is released from the plasma membrane and accumulates in the nucleus to activate gene expression for hypoxia acclimation. Our discovery of an oxygen-sensing mechanism opens up new possibilities for improving flooding tolerance in crops.
大多数的真核生物依赖于分子氧用于呼吸能量的产生。当供氧不足时,机体会激活一系列的适应性反应来减少能量消耗带来的有害性影响。已经有很多种诱导这类反应的氧气感应机制被报道,但他们似乎都是各类生物特有的,并且植物中还未曾有过相关方面的报道。本文中,作者发现了一类在动植物中均作用于蛋白降解的泛素依赖性N端规则通路在拟南芥中作为氧气感应机制发挥功能。作者鉴定了乙烯响应因子RAP2.12氨基酸序列的保守末端氨基酸作为一个翻译后修饰的氧气依赖性序列,最终会导致有氧条件下RAP2.12的降解。当水淹情况下氧气浓度较低时,RAP2.12从质膜上被释放,并且在核中积累,已激活一些缺氧适应相关基因的表达。本文的发现揭示了植物中的一个氧气感应机制,为提升作物的抗洪涝灾害能力提供了一个新的可能。
通讯:Joost T. van Dongen (https://www.mpimp-golm.mpg.de/11030/Joost_van_Dongen)
个人简介:1997年,荷兰乌得勒支大学,学士;2001年,荷兰乌得勒支大学,博士。
研究方向:植物对低氧胁迫的生物和分子响应。
doi: https://doi.org/10.1038/nature10536
Journal: Nature
Published date: November 17, 2011
Homeostatic response to hypoxia is regulated by the N-end rule pathway in plants
First author: Daniel J. Gibbs; Affiliations: University of Nottingham (诺丁汉大学): Loughborough, UK
Corresponding author: Michael J. Holdsworth
Plants and animals are obligate aerobes, requiring oxygen for mitochondrial respiration and energy production. In plants, an unanticipated decline in oxygen availability (hypoxia), as caused by roots becoming waterlogged or foliage submergence, triggers changes in gene transcription and messenger RNA translation that promote anaerobic metabolism (无氧代谢) and thus sustain substrate-level ATP production. In contrast to animals, oxygen sensing has not been ascribed to a mechanism of gene regulation in response to oxygen deprivation in plants. Here we show that the N-end rule pathway of targeted proteolysis acts as a homeostatic sensor of severe low oxygen levels in Arabidopsis, through its regulation of key hypoxia-response transcription factors. We found that plants lacking components of the N-end rule pathway constitutively express core hypoxia-response genes and are more tolerant of hypoxic stress. We identify the hypoxia-associated ethylene response factor group VII transcription factors of Arabidopsis as substrates of this pathway. Regulation of these proteins by the N-end rule pathway occurs through a characteristic conserved motif at the amino terminus initiating with Met-Cys. Enhanced stability of one of these proteins, HRE2, under low oxygen conditions improves hypoxia survival and reveals a molecular mechanism for oxygen sensing in plants via the evolutionarily conserved N-end rule pathway. SUB1A-1, a major determinant of submergence tolerance in rice, was shown not to be a substrate for the N-end rule pathway despite containing the N-terminal motif, indicating that it is uncoupled from N-end rule pathway regulation, and that enhanced stability may relate to the superior tolerance of Sub1 rice varieties to multiple abiotic stresses.
植物和动物都是专性需氧型生物,需要氧气用于线粒体呼吸和产生能量。在植物中,根或叶片遭水淹导致植物组织可用氧气含量的突然下降,进而诱导基因转录和信使RNA翻译的改变,从而促进无氧代谢,最终维持底物水平ATP的生产。与动物相反,植物中响应于缺氧所诱导的基因表达调控尚未能归因于某一个特定的分子机制。本文中,作者发现靶向蛋白酶解的N端规则通过调控关键缺氧反应转录因子,作为拟南芥面临极度低氧水平的一个稳态感应器发挥作用。作者发现植物缺少N端规则途径组分会导致核心缺氧反应基因的组成型表达,并且对低氧胁迫的耐受性更高。作者发现拟南芥中缺氧相关的乙烯响应因子VII类转录因子是该途径的底物。N端规则途径通过一个保守的氨基酸序列末端Met-Cys特征性基序来调节这些蛋白质。通过增强这些蛋白质之一HRE2在低氧条件下的稳定性,提高了低氧存活率,说明植物通过进化上保守的N端规则途径感应氧气浓度的分子机制。SUB1A-1是水稻水淹耐受性的主要决定因子,而该蛋白尽管含有保守的N端基序,但并不是N端规则途径的底物,这表明它与N端调控途径没有关联,并且增强的稳定性可能与Sub1水稻品种对多种非生物胁迫的超强耐受性有关。
通讯:Michael J. Holdsworth (https://www.nottingham.ac.uk/biosciences/people/michael.holdsworthv)
个人简介:1999年,瑞士苏黎世联邦理工学院,学士;2003年,瑞士苏黎世联邦理工学院,博士;2003-2009年,德国弗莱堡大学,博士后。
研究方向:植物N-degron通路在生长、发育以及对环境响应等方面的作用。
doi: https://doi.org/10.1038/nature10534
Journal: Nature
Published date: November 17, 2011
P.S. 感兴趣的还有一篇综述可以参考参考,链接:https://doi.org/10.1146/annurev-arplant-043014-114813(下图来自该文)
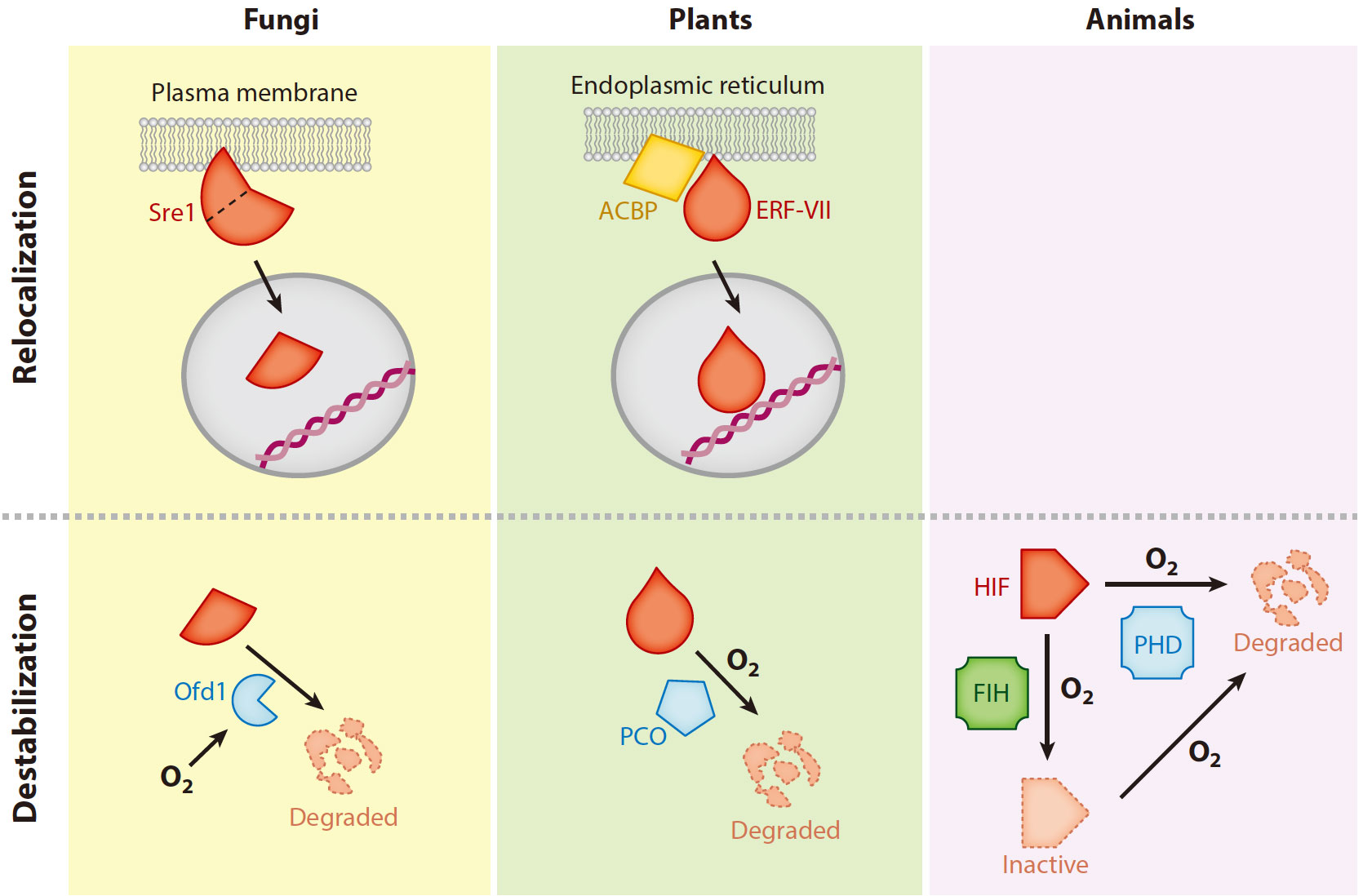
Figure 4. Cross-kingdom comparison of low-oxygen-sensing mechanisms. Oxygen-dependent relocalization of a transcription factor that acts as a master regulator of the anaerobic response (shown in red ) from membrane systems into the nucleus is common to fungi and plants. In Schizosaccharomyces pombe, the transcription factor Sre1 is cleaved in an indirect oxygen-dependent manner, thereby releasing an active N-terminal fragment into the nucleus. In Arabidopsis thaliana, RAP2.12 is localized at the plasma membrane, where it interacts with acyl–coenzyme A (CoA)–binding proteins (ACBPs). Hypoxic conditions trigger its release by an unknown mechanism. Oxygen-dependent destabilization of the hypoxic master regulator is also shared among eukaryotes, although through different mechanisms: In yeast, Sre1 is degraded by the protease Ofd1, whose activity is directly stimulated by oxygen binding. In plants, plant cysteine oxidases (PCOs) prepare RAP2.12 for protein degradation by oxidizing the N-terminal cysteine. In animals, the HIF-1α transcription factor is hydroxylated at a proline residue in the presence of oxygen by prolyl dehydrogenases (PHDs), a modification that initiates HIF degradation by the proteasome. A second hydroxylation at an asparagine residue by FIH also inactivates the transcription factor.

https://blog.sciencenet.cn/blog-3158122-1201249.html
上一篇:New Phytologist:植物中光信号转导的演化
下一篇:Current Biology:植物根-茎系统性信号转导介导根结线虫抗性
全部作者的其他最新博文
- • Plant Physiology:CsMADS3促进柑果中的叶绿素降解和类胡萝卜素合成(华中农业大学)
- • Molecular Plant:LBD11-ROS反馈调节作用于拟南芥的维管形成层增殖和次生生长(浦项科技大学)
- • Science Advances:根结线虫通过调控植物的CLE3-CLV1模块,促进侵染进程(日本熊本大学)
- • Nature Communications:油菜素内酯参与植物营养生长期转变的分子机制解析(浙江农林大学)
- • Current Biology:光合作用产生的蔗糖驱动侧根“生物钟”(德国弗莱堡大学)
- • PNAS:花同源异型基因在叶中被抑制、花中被激活的分子机制(南卡罗来纳大学)