博文
组学能否定量? 组学如何定量?
|
谨慎使用“定量组学”
准确定量是分析化学的必要条件,也是脂质组学与代谢组学可持续发展所不可或缺的重要内容,从而使脂质组学与代谢组学在生物和生物医学领域广泛应用。
以脂质组学为例,为了保证数据准确量化,需要在整个脂质组学工作流程中满足特定的条件(图1)。多年来,脂质分析已经发生了巨大的变化,从运用只允许检测一组或几组相当有限的脂质种类/类别的传统技术到能分析数量能达到数百甚至数千种呈指数性爆发的生物脂质组(图2)。质谱技术的出现和进步是这些变化的主要催化剂,随着方法学的不断改进,加上准确性、特异性、敏感性以及可检测的脂质种类数量的提高,脂质组学正稳步从比较脂质组学向准确的定量组学过渡。近年来,随着脂质组学研究的大量涌现,为了保证脂质组学数据准确定量,也许是时候让研究人员为其制定适当的术语定义和分析考虑因素,以便使其能应用于更大的临床领域(例如疾病生物标志物阈值的定义)。实际上,从长远来看,定量的准确性可能会对脂质组学及代谢组学的持续发展产生重大影响。
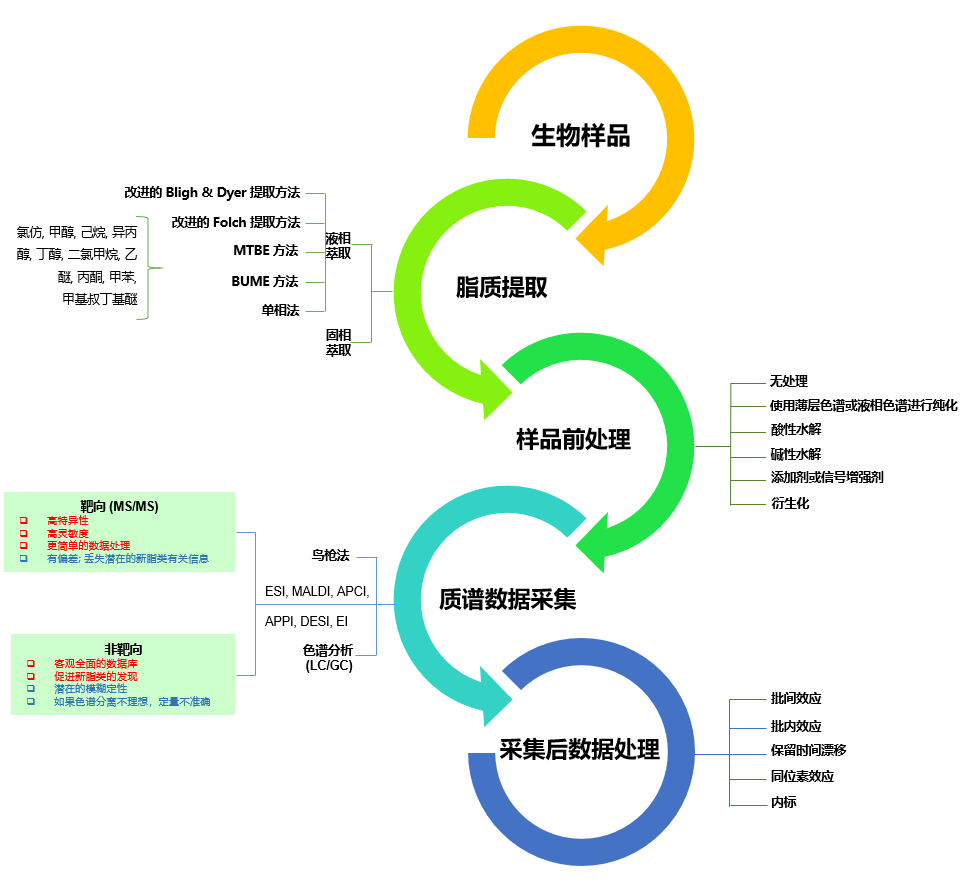
图1.影响典型脂质组学工作流程中定量准确性的因素。常规工作流程中的各种因素可能引入也许会影响最终定量准确性的错误源。
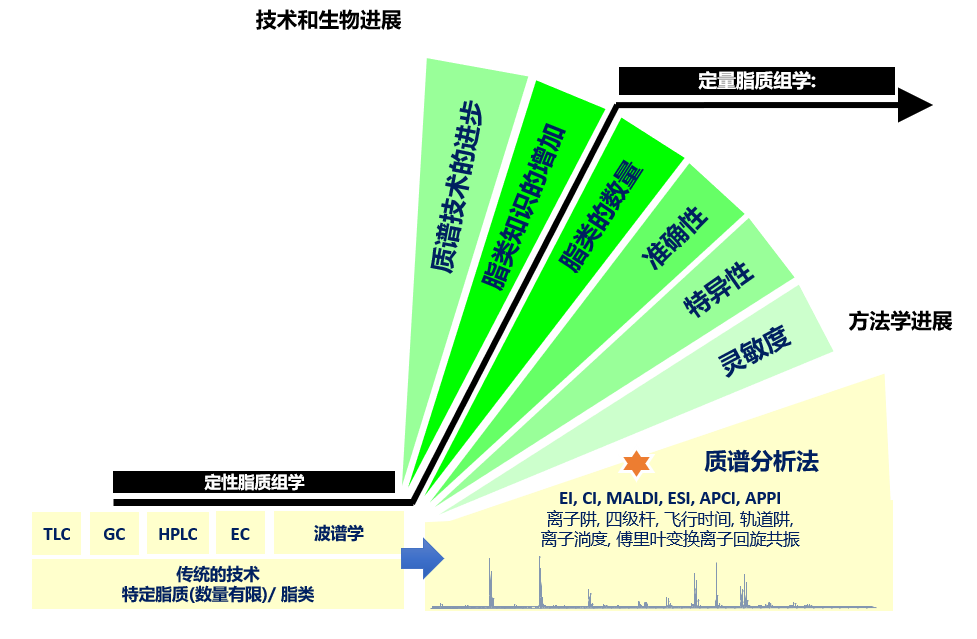
图2.脂质组学向定量准确性的转变。质谱技术的进步推动了脂质组学方法的开速发展,朝着定量准确的最终目标迈进,这是脂质组学作为组学科学新兴分支不断发展壮大的关键决定因素。
在质谱分析中严格要求使用同位素标记的内标来实现绝对定量是一个公认的事实,这主要是由于任何分子离子的离子计数(即强度)与其绝对浓度之间缺乏明确的联系。基于这一严格的定义,在没有与被分析物质具有相同电离和破碎特性的同位素标记类似物的情况下,任何采集后数据校正都只是对不严谨定量分析方法的修正以提高量化分析的准确度,都不能实现准确的绝对定量。由于生物的主要代谢物缺少市售的同位素标记类似物,再加上对生物代谢组的每一种代谢物一对一定制合成同位素标记内标所需的高昂成本,使得绝对定量在实际应用中过于理想化而不现实。相对于绝对定量,高精度的相对定量通常足以满足当前大量的以组学为导向的科学和转化研究的要求。然而,为了获得准确的定量,内标的性质和数量需要依次满足相关的具体要求。
通常,含有奇数链脂肪酸或稳定同位素标记类似物的脂质被加入样品中作为定量的内标。使用奇数链脂肪酸组成的脂作为内标是基于这些脂肪酰通常在高等生物(例如哺乳动物)中的内源性丰度较低(通常<1%)的初步检测;但是,对不同的组织如大脑则需要采取谨慎的态度,因为大脑中可能含有比较高比例的奇链脂肪酸[1]。此外,奇数链脂肪酸的作为内标也不适用于植物、微生物和秀丽隐杆线虫等模式生物,因为这些模式生物本身含有大量奇数链脂肪酸[2]。因此,没有简单的规则来证明奇数链脂肪酸内标的应用是合理的,必须提前确定了确切的模式生物和组织/细胞类型,排除了奇数链脂肪酸的存在,然后才能应用相应的奇数链脂肪酸内标。这突出了为脂质研究中常用的每种模式生物构建一个全面的脂质库的重要性[3],因为这些有价值的信息可以为研究人员在不同研究中选择内标时提供便利。另一方面,稳定同位素标记的类似物是精确定量的理想内标,因为它们相同的化学性质将产生与对应的内源物质相同的离子响应因子。然而,建议在使用同位素标记的标准品前先验证它们的纯度,因为稳定同位素标记中的杂质可能会导致与相邻内源性物质的混淆峰,特别是在没有HRMS的情况下,会导致定量误差。
对于标准品的数量,为了校正每种脂质类别中各个类别的不同酰基链长度所导致的离子响应因子的差异,通常推荐使用两个或两个以上的与分析脂类相对应的内标。这对中性脂质如三酰甘油(TAG)和蜡酯(WES)尤其重要[4],由于其缺少极性头部基团,导致脂肪酰链长度和不饱和度对脂质的整体极性的贡献更大,由此影响离子对质谱的响应[5]。此外,由于中性脂质作为能量储存和/或细胞外结构成分发挥着主要作用[3] ,它们通常比极性脂质在脂肪酰链长度的范围更大。因此,为了准确地定量每类中性脂质,有时可能需要三个或三个以上跨酰基链长度有效范围的内标。
除了目标脂质分子的整体极性外,使用MS/MS进行精确定量时要考虑的另一个关键问题是目标化合物的裂解模式相对于作为标准化的内标的比较。MS/MS的使用通过降低基线漂移和背景噪声大大扩展了分析方法的线性动态范围,这主要归功于串联质谱提供的双重过滤。然而,必须注意的是,在碰撞诱导的解离过程中,同一种脂质内的不同类别的裂解可能会发生变化,这主要是由于解离动力学的不同和产物碎片的热力学稳定性不同造成的。因此,在定量时需要谨慎选择内标,以减少串联质谱因子引入的变化[5]。这对于单个的MRMs不是建立在丢失一个共同碎片,而是建立在不同的碎片(例如,连接在每类脂质上的单个脂肪酰基)的脂类尤其相关。
-部分内容摘自脂质组学专家评述专刊封面论文“Lipidomics,en route to accurate quantitation”,BBA Mol Cell Biol Lipidomics,2017,1862(8):752-761.
Defining “Quantitation” in Omics Research
Accurate quantitation is the sine qua non of analytical chemistry, and is indispensable for the sustainable development of lipidomics via enabling its wide applications in the biological as well as biomedical settings. Specific conditions needs to be fulfilled along the entire lipidomics workflow in order to qualify the data as accurately quantitative (Figure 1). Over the years, lipid analysis has evolved tremendously, from the application of traditional techniques that only permit the detection of a rather limited set of lipid species/classes to an exponential outburst in the sizes of the characterized biological lipidomes, often culminating to hundreds or even thousands of individual species in number (Figure 2). The emergence and advancement in MS technologies is the major catalyst for these changes, and with the continual improvements in methodology, coupled with the accompanying augmentations in accuracy, specificity, sensitivity as well as the number of detectable lipid species, lipidomics is transiting steadily from being solely qualitative towards accurate quantitation. With the outpour in lipidomics studies over the recent years, perhaps it is timely for researchers to lay out proper definitions of terms and analytical considerations for lipidomics data to be deemed sufficiently quantitative in order to render their applications in the larger clinical settings (e.g. in the definition of disease biomarker thresholds). In essence, quantitative accuracy may substantially impact upon the persevering development of lipidomics in the long run.
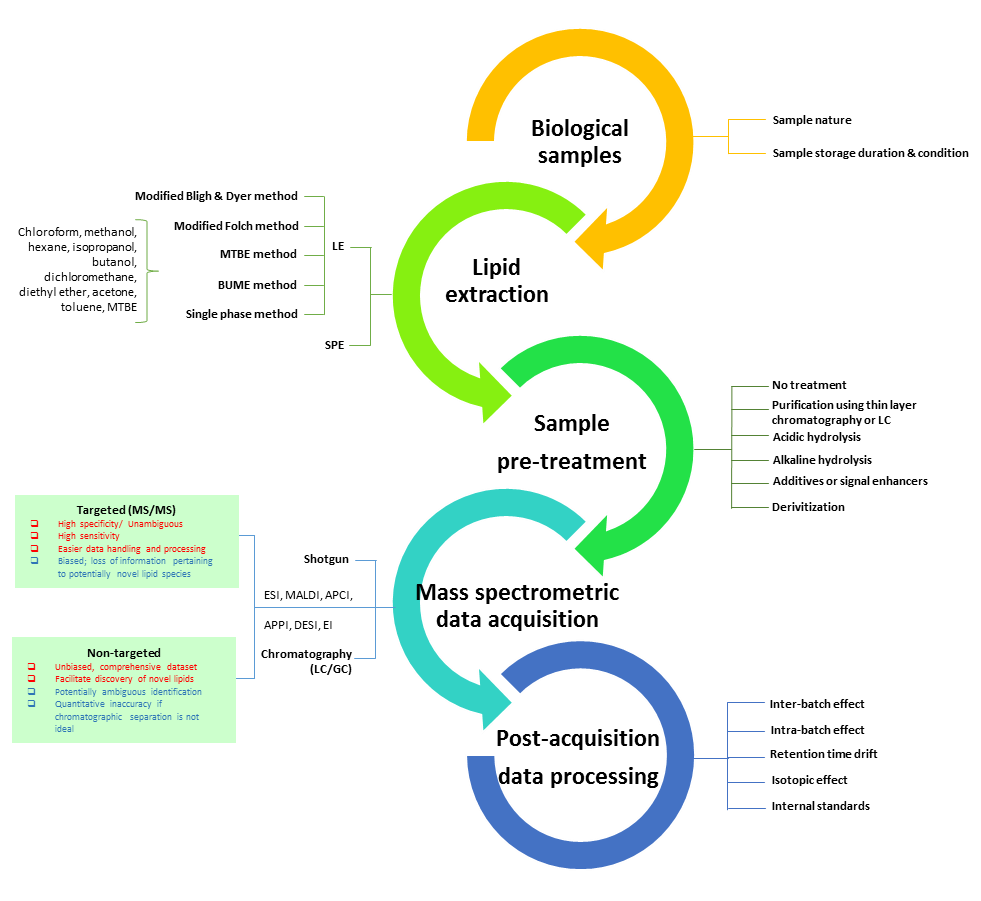
Figure 1. Factors that may impact upon quantitation accuracy along a typical lipidomics workflow. Various factors along the routine workflow can potentially introduce sources of errors that may compromise the ultimate quantitative accuracy.
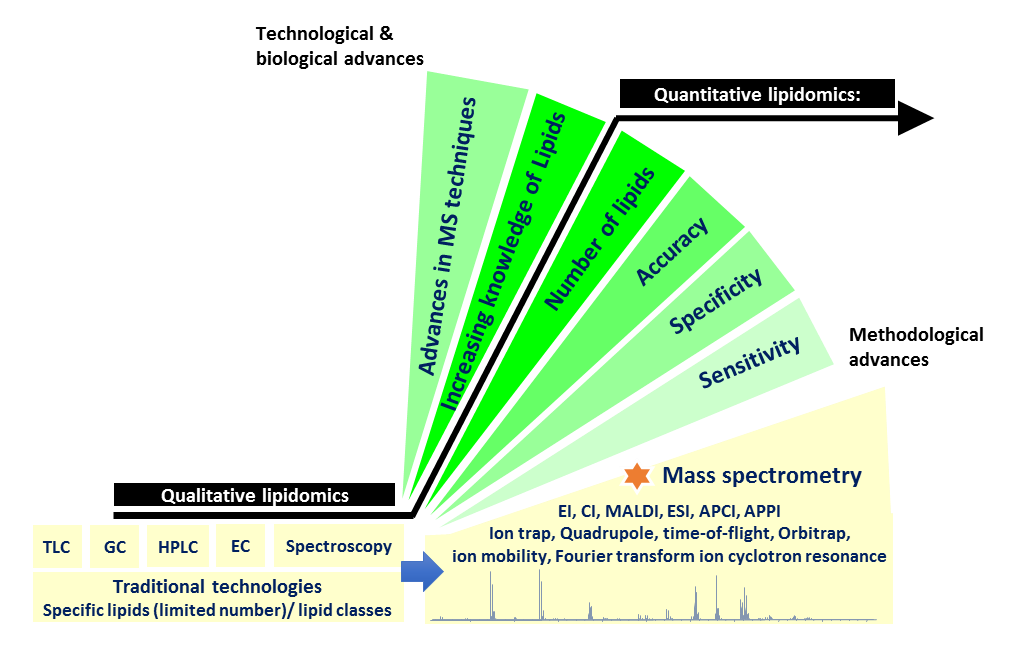
Figure 2. the transformation of lipidomics towards quantitative accuracy. Advancements in mass spectrometry has fueled the rapid methodological developments in lipidomics towards the ultimate goal of quantitative accuracy, a key determinant towards the continual growth and expansion of lipidomics as an emerging branch of omics science.
The strict requirement for isotopically-labelled internal standards to achieve absolute quantitation on the MS is a recognized fact, which is principally attributed to the lack of a definite association between ion counts (i.e. intensity) of any molecular ion with its absolute concentration. Based on this strict definition, in the absence of isotope-labelled analogs with identical ionization and fragmentation properties to the analyte itself, no amount of post-acquisition data correction will achieve absolute quantitation. The lack of commercially available isotope-labeled analogs for a major portion of the biological metabolome, coupled with the soaring costs for one to custom-synthesize isotope-labelled standards for each and every single metabolites constituting the biological metabolome, have made absolute quantitation more idealistic than realistic to attain in real life applications. Instead of absolute quantitation, relative quantitation with high accuracy is often sufficient to fulfill the requirements for a prevailing amount of omics-oriented scientific and translational studies. Nonetheless, specific requirements pertaining to the nature and number of internal standards need to be satisfied in order to attain accurate quantitation.
Typically, lipids containing odd-chain fatty acids or stable isotope-labelled analogs are spiked into the samples as internal standards for quantitation. The use of comparable lipids comprising odd-chain fatty acids is based on preliminary observations that such fatty acyls are usually found in low endogenous abundance (typically <1 %) in higher organisms (e.g. mammals); but caution needs to be exercised for distinct tissues such as the brain, which may contain notably higher proportion of such odd-chain fatty acids [1]. Furthermore, standards based on odd-chain fatty acids will not be applicable for organism models such as plants, microbes and Caenorhabditis elegans, which inherently comprise appreciable amounts of odd-chain fatty acids [2]. Thus, there is no simple rule to justify the application of such internal standards, and the conditions for distinct organismal models and tissue/cell types must be pre-determined to exclude the presence of odd-chain acids before the corresponding standards can be applied. This highlights the critical importance of constructing an all-encompassing lipidome inventory for each model organism commonly utilized in lipid research [3], since these valuable information can come in handy for researchers in their choice of internal standards for different studies. On another note, stable isotope-labeled analogs denote ideal internal standards for accurate quantitation, as their identical chemical properties would translate to equal ion response factors to the corresponding endogenous species. Nevertheless, it is advisable to first verify the purity of these isotopically-labeled standards prior to use, as impurities in stable isotope-labelling may potentially lead to confounding peaks with adjacent endogenous species, especially in the absence of HRMS, resulting in quantitative errors.
As for the number of standards, in order to correct the differences in ion response factors attributed to varying acyl chain lengths for individual species within each lipid class, two or more internal standards corresponding to each class of lipids being analysed is usually recommended. This is particularly important for neutral lipids such as the triacylglycerols (TAGs) and wax esters (WEs) [4], since the absence of polar head groups would translate greater contribution of fatty acyl chain lengths and unsaturation to the overall polarity of individual species, and henceforth the resulting ion response on MS [5]. Furthermore, neutral lipids often span a greater range than polar lipids in terms of fatty acyls chain lengths, given their predominant roles as energy stores and/or extracellular structural components [3]. As such, three or more internal standards spanning an effectively wide range in acyl chain lengths may at times be required for accurate quantitation of each class of neutral lipids.
Besides the overall polarity of the lipid molecules of interest, another critical issue to consider in accurate quantitation using MS/MS is the fragmentation pattern of the compound of interest relative to the internal standard for which normalization is based upon. The use of MS/MS greatly extends the linear dynamic range of an analytical method by virtue of its reduced baseline drift and lowered background noise, primarily attributed to double filtering offered by tandem MS. It must be noted, however, that fragmentation of individual species within the same lipid class during collision induced dissociation can be varied, principally due to the distinct kinetics of dissociation and the differing thermodynamic stability of resultant product fragments. As a result, care needs to be exercised in the selection of internal standards during quantitation in order to curtail variations introduced by the tandem MS factor [5]. This is especially relevant for lipid classes for which MRMs for individual species are not constructed upon the loss of a common fragment, but distinct fragments instead (e.g. individual fatty acyls attached to each species).
- Excerpt edited from “Lipidomics, en route to accurate quantitation”, cover paper for special issue on lipidomics opinion articles from BBA Mol Cell Biol lipids, 2017, 1862(8): 752-761
参考文献:
1. S.M. Lam, G.H. Chua, X.J. Li, B. Su, G. Shui, Biological relevance of fatty acyl heterogeneity to the neural membrane dynamics of Rhesus macaques during normative aging, Oncotarget (2016).
2. S.M. Lam, Z. Wang, J. Li, X. Huang, G. Shui. Sequestration of polyunsaturated fatty acids in membrane phospholipids of Caenorhabditis elegans dauer larva attenuates eicosanoid biosynthesis for prolonged survival, Redox Biology (2017).
3. S.M. Lam, G. Shui, Lipidomics as a principal tool for advancing biomedical research, J Genet Genomics 40 (2013) 375-390.
4. S.M. Lam, L. Tong, B. Reux, M.J. Lear, M.R. Wenk, G. Shui, Rapid and sensitive profiling of tear wax ester species using high performance liquid chromatography coupled with tandem mass spectrometry, J Chromatogr A 1308 (2013) 166-171.
5. M. Wang, C. Wang, X. Han, Selection of internal standards for accurate quantification of complex lipid species in biological extracts by electrospray ionization mass spectrometry-What, how and why?, Mass Spectrom Rev (2016).
Sin Man Lam 博士简历
中科脂典技术总监。2009年获得新加坡国立大学生物学一级荣誉学士学位,同年获新加坡国立大学华莱士奖项(授予生物系第一名毕业生),新加坡国立大学校长研究生奖学金(President Graduate Fellowship)获得者。2013-2017年,在中国科学院遗传与发育生物学研究所工作; 2014年获益海嘉里优秀博士后奖项, 同年获得第7届中国博士后特别资助; 2015年获国家自然基金委青年基金资助。研究方向包括基于液相色谱与质谱联用的高通量、高覆盖脂质组和代谢组半定量/精准定量方法开发,并应用于临床和生物学研究中。博士期间建立了最全面的泪液、睑板腺脂质谱图并应用于干眼病治疗过程的检测分析,发现治疗干眼病潜在新靶点(系列文章发表在Journal of Lipid Research, 3篇)。自2010年来,在Science, Cell Research, Development Cell, PNAS, Redox Biology, Nature Communications, Cell Reports, Diabetes, Plos Pathogens, Plant Physiology, Plos Genetics, Neurobiology of Aging, Journal of Lipid Research, BBA-molecular and cell Biology of Lipids, Analytica Acta Chimica, Journal of Chromatography A, Metabolomics等国际期刊发表41篇SCI文章。
(本文由Sin Man Lam和税光厚共同完成)
https://blog.sciencenet.cn/blog-3356615-1190261.html
上一篇:2019脂质代谢调控盛会:5th Lipidall会议第一轮通知
下一篇:第五届LipidALL脂质代谢会议第2轮通知(内容丰富,阵容强大!)